A Nobel Recognizes a Cool Scientific Method (Pun Intended)
 Joachim Frank. Photo Jörg Meyer.
Joachim Frank. Photo Jörg Meyer.Until late last year, an amber-colored structure stood on a window sill in Joachim Frank’s lab, where it glowed, backlit by the late morning sun. You would be forgiven for thinking that this series of oddly shaped plastic contours, serially mounted in a wooden frame, was a topographical map of some kind. But the object was a 3-dimensional reconstruction of a part of a molecular machine in the cell called a ribosome, the first reconstruction of its kind and a creation made in Dr. Frank’s lab more than three decades ago.
In December, Dr. Frank packaged it up and shipped it to the Nobel Museum in Stockholm, where it now lives as an artifact representing his journey toward the 2017 Nobel Prize in Chemistry. Dr. Frank, a physicist-turned-structural biologist, was one of the trio of scientists awarded the prize. He and his co-laureates, Jacques Dubochet and Richard Henderson, were honored for their work in developing a technique called cryo-electron microscopy (cryo-EM) that today allows researchers to visualize biological molecules—proteins, RNA, and DNA—in exquisite detail.
 © Nobel Media AB 2017 Photo / Alexander Mahmoud
© Nobel Media AB 2017 Photo / Alexander Mahmoud“A cool method for imaging the molecules of life” was how Professor Göran K. Hansson, secretary general of the Royal Swedish Academy of Sciences, described the technique in announcing the prize on the morning of Oct. 4. His pun referred to a key step in cryo-EM: suspending the sample in water rapidly cooled to form a glasslike structure that acts as a freeze-frame and a contrast enhancer for biological molecules.
Dr. Frank, professor of biochemistry & molecular biophysics at VP&S and professor of biological sciences at Morningside, was not the refrigeration guy; he was the image guy. His contribution to the technique over his 40-plus-year career has been to develop powerful computational and statistical methods that combine multiple blurry images into one sharp one. For years, structural biologists dismissed electron microscopy as “blobology,” convinced that it would never yield high-resolution structures. But Dr. Frank’s approach, called single particle reconstruction, helped set the stage for a molecular imaging revolution, creating a technique that can capture every bend, twist, fold, and pocket in a protein’s structure to provide an unprecedented view of how these molecules conduct their cellular business. Along the way, Dr. Frank’s lab—based at VP&S for the past decade—has revealed some fundamental elements of how a cellular machine called the ribosome does its crucial job of synthesizing proteins.
“It is incredibly gratifying to see this kind of acknowledgment of the whole field,” says Dr. Frank.
Attracted to Science at an Early Age
Seated in his VP&S lab’s conference room just two days after returning from the Nobel ceremony in Stockholm, Dr. Frank, dressed casually in blue trousers, shirt, sweater, and a well-worn pair of sneakers, initially seems a bit exhausted by the prospect of giving yet another interview about his life’s work. But it is not long before the gleam returns to his eyes as he reaches deep into the history of structural biology and of his own research.
Dr. Frank was drawn to science at an early age, an interest he attributes to a search for order spurred by the tumultuousness of his wartime childhood in Siegerland, Germany. He first bumped shoulders with electron microscopy as a master’s student in physics in the mid-1960s: His mentorhad an old electron microscope in his office that he fiddled with, and the technique piqued Dr. Frank’s interest. Poking around in physics abstracts, he found a researcher named Walter Hoppe at the Technical University of Munich who wanted to use electron microscopy to image biological molecules. Dr. Frank showed up at his door and signed on for a PhD.
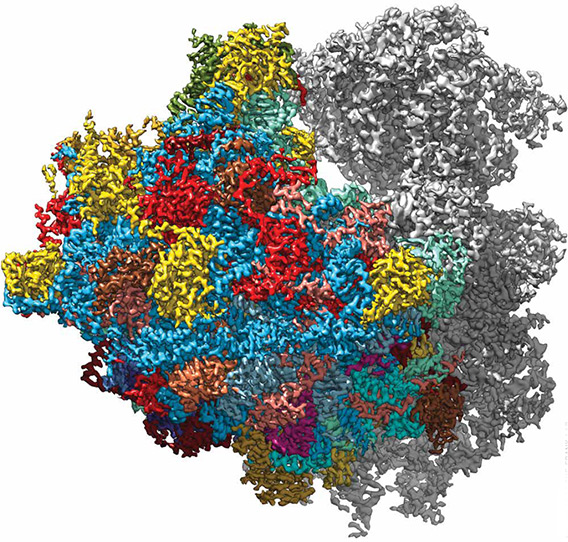 Reconstruction of the Trypanosoma cruzi ribosome at 2.5 angstrom resolution, obtained in Dr. Frank’s lab. The resolution in large parts of the molecule should be sufficient for design of effective drugs against Chagas disease. Illustration courtesy of the Frank lab.
Reconstruction of the Trypanosoma cruzi ribosome at 2.5 angstrom resolution, obtained in Dr. Frank’s lab. The resolution in large parts of the molecule should be sufficient for design of effective drugs against Chagas disease. Illustration courtesy of the Frank lab.Electron microscopy was invented in the 1930s, but initially no one envisioned using it for biology because it requires the sample to be placed in a vacuum—in which biological molecules collapse into a structureless heap—and because the image is achieved by bombarding the sample with electrons, which also destroys its structural integrity. Dr. Hoppe had started out using a technique called X-ray crystallography, which remains a mainstay of structural biology. However, not all molecules can be crystalized, and for those that can, the crystal structure might very well differ from the molecule’s native state. “The crystal formation selects one conformation to the exclusion of others,” Dr. Frank explains. “And the conformation that’s selected might not be functionally relevant.”
When Dr. Frank started his doctorate, he found himself at the heady center of structural biology’s early days as Dr. Hoppe and a handful of other pioneers set out to use electron microscopy to get around these problems. By the time he defended his dissertation in 1970, computer-based image processing had advanced to the point where researchers could envision enhancing images obtained with low-dose electron microscopy.
The generally accepted approach was to tilt the copper grid on which the sample lay inside the microscope, capturing the projection of the molecule serially at each angle and then mathematically combining the series of resulting micrographs into a single image. But even at low doses, walloping a sample with electron beams multiple times destroyed it. What’s more, the approach worked primarily for symmetrical or regularly ordered molecules. Generalizing the technique to tackle asymmetry was crucial because most molecules in the cell, and complexes of molecules that form to perform specific cellular functions, are not symmetrical.
Dr. Frank thought he had a better idea. After all, an electron microscopy sample consists of hundreds of thousands of copies of a biological molecule, each copy in a random orientation. Rather than repeatedly beaming the same molecule multiple times, one could take a snapshot containing multiple copies of the molecule. It should be possible to work backward to determine their viewing angles to reconstruct them. He described the idea in a concept paper in 1975, but it did not make much of a splash. “The community thought I was crazy, that it was never going to work,” recalls Dr. Frank.
That same year, Dr. Frank took up his first independent position at the Wadsworth Center, a public health research facility in Albany, N.Y. He began trying to realize his theoretical idea with various molecules, knowing that the big difficulty would be to develop a computational method for averaging traces of molecules that were captured in different conformations or from different views. Back then, researchers ran their analyses in shared computing centers, on computers as big as refrigerators that packed about a thousandth of the computational punch of an iPhone, he recalls. But on the plus side, these facilities were a venue in which scientists from different fields could interact. In 1980, a lucky glance one afternoon at data printouts belonging to a laboratory medicine researcher gave him the jolt of insight he needed to design a powerful statistical framework to solve the problem.
Meanwhile, another chance encounter introduced Dr. Frank to the ribosome, which became his lab’s workhorse and muse. Ribosomes are essentially mobile protein-building machines: They translate the instructions of the DNA code, using the messenger RNA code to string together amino acids carried by transfer RNA. Yet the molecular mechanics of the process were largely unknown. Because of the organelle’s complexity, size, and asymmetry, Dr. Frank immediately embraced it as the perfect test specimen.
 Members of the Frank lab. Photo Jörg Meyer.
Members of the Frank lab. Photo Jörg Meyer.Fueled by these developments, the lab was a tremendously productive place to work, says Jose-Maria Carazo, PhD, now head of the biocomputing unit at the National Center for Biotechnology in Madrid, who arrived as a postdoc in 1986. “Joachim had everything in place so we could advance quickly,” Dr. Carazo says. “His philosophy was that you should not try to acquire tools from here and there; rather, he wanted to create an enabling environment.”
The ribosome became his lab’s workhorse and muse. Because of its complexity, size, and asymmetry, Dr. Frank immediately embraced it as the perfect test specimen.
That year, Dr. Frank and a postdoc, Michael Radermacher, succeeded in creating the first 3-dimensional reconstruction of the larger subunit of the ribosome—the very one that Dr. Frank sent to the Nobel Museum. A few years later came a structure of the whole molecule, though its resolution, at 45 angstroms, was more than an order of magnitude larger than the atomic scale that X-ray crystallography can achieve. Still, it captured the attention of a few biologists, among them Rajendra K. Agrawal, then a PhD student in biochemistry at Bananas Hindu University in India, who happened to be studying the ribosome. Dr. Agrawal’s research was based on the theory that this messenger RNA-reading machine functioned by switching between at least two conformations, and he thought Dr. Frank’s method could prove it.
“To tell you the truth I was sort of naïve about the resolution limits of the technique at the time,” says Dr. Agrawal, who went to the Frank lab for a postdoc in 1994. “When I saw that paper, I got excited without paying attention to the resolution, which was very, very low.” He arrived to a lab filled mostly with computers and microscopes, with just a tiny space for the kinds of biochemistry experiments he was accustomed to doing. Most of the team was developing the 3D-EM method, not studying biology. “It was kind of a gentle shock,” he recalls. “For a month or so, I thought, ‘I hope I didn’t make a mistake!’”
It turned out to be no mistake at all. His first paper in the lab, published within a year of his arrival, brought the resolution down to 25 angstroms, a point at which the molecule began to yield some big functional mysteries. This reconstruction allowed the researchers to visualize how the ribosome interacted with mRNA and tRNA as it went about the process of translation. For Dr. Frank, that paper was a turning point. “Finally, I thought, ‘Oh my God, I can really make contributions to biology,’” he says. “Before that it was not so clear.”
Dr. Frank’s lab continued to improve the resolution of the technique, making several important discoveries along the way at a rapid clip. The researchers were able to see where, exactly, the tRNA makes contact with the ribosome and shortly thereafter to visualize a protein called the elongation factor G binding to the ribosome to advance mRNA and tRNA forward, thereby building the amino acid chain. They also managed to accomplish what Dr. Agrawal had most hoped to achieve: clearly seeing the two conformations of the ribosome.
At the same time, another research team was homing in on an atomic-resolution structure using X-ray crystallography. “I remember we were coming back to Albany on the same flight, and I found him a little dejected because he thought it looked like it was all over for us,” says Dr. Agrawal. “But I said, ‘Joachim, I think it’s all only starting for us.’” X-ray crystallography could capture only fixed states of a molecule, but cryo-EM could capture the dynamics. By going back and forth between the two techniques, a fuller picture of the protein’s function could be built. And that’s just how it happened: In 2000, Dr. Agrawal and colleagues published a paper demonstrating the ratcheting motion that the two subunits make as the ribosome moves down the mRNA strand. “That was the first time that a real, very large conformational change was discovered,” Dr. Frank recalls. “People got excited by the fact that you could really see molecules in multiple conformations and you could really see they differed from what actually crystallography told you.”
Warm and Nurturing Environment
Despite the intensity of the work in the Frank lab, trainees who have passed through the lab remember it as a warm and nurturing environment and Dr. Frank as a mentor with a light touch, a deep sense of humor, and a few subtle surprises up his sleeve. Robert Grassucci, the lab’s senior microscopist, came on board in 1986 as a technician. He was in his freewheeling 20s at the time, and his much older boss initially seemed somewhat staid, the stereotype of a German scientist. Mr. Grassucci stopped by the lab one weekend and found Dr. Frank playing some unexpected music. “I was like, ‘Huh, that’s kind of cool, what’s this old prof doing jamming to Bob Marley?’” he recalls. “So I thought, OK, that’s something we have in common.”
Many of Dr. Frank’s students and postdocs were from overseas, and he and his wife, Carol Saginaw, took them in for holidays and made sure they settled in and found their footing. “I was not paying attention to those things then, but when I think back now, he was such a family-oriented person all along, always trying to make us comfortable,” Dr. Agrawal says, noting that Dr. Frank would often point him to Indian restaurants in town or drive him to see music concerts.
"People got excited by the fact that you could really see molecules in multiple conformations."
By that point, the group had brought the resolution of cryo-EM down to 11.5 angstroms, making it possible to see the double helix of the RNA and even some features of proteins in the ribosome. Dr. Frank and his team continued probing the mechanism of this molecule in action. In 2001, his postdoc Christian Spahn, now a professor at the Charite Institute of Medical Physics and Biophysics in Berlin, collaborated with Jennifer Doudna’s lab, then at Yale, to create a cryo-EM image of the virus hepatitis C hijacking a cell’s DNA via its ribosome with the help of an RNA element called an internal ribosome entry site. “We could see this element grabbing and holding on to the small subunit of the ribosome, which was really spectacular,” says Dr. Frank. “There are these rare moments where you look at a structure and suddenly realize that there is really something fascinating—and that you are probably the first human being to understand this particular thing,” says Dr. Spahn, who was a postdoc in the Frank lab between 1998 and 2004.
Ultimately, it was not cryo-EM that first pinned down the structure of the ribosome at atomic scale but X-ray crystallography. That achievement was recognized with the 2009 Nobel Prize in Chemistry, and Dr. Frank’s work was fundamental to the work of one of the recipients who used a reconstruction produced in the Frank lab to check the validity of the structure. It took until the advent of a new type of electron-detecting camera in 2012 for researchers to achieve the atomic structure of the ribosome using cryo-EM. That technology, Dr. Frank explains, has democratized the method. “The technique was sort of waiting for that breakthrough,” he says. Indeed, in 2015 cryo-EM was named Nature magazine’s Method of the Year.
“Joachim didn’t just shape the field directly, with his intellectual contributions of the math and physics, but also indirectly,” says Yaser Hashem, PhD, group leader at the Institute of Cellular and Molecular Biology in Strasbourg. “By using the technology and constantly being in contact with the companies that made it, his work really guided the development of the microscopes.”
The Other Joachim Frank
In parallel to research, Dr. Frank actively pursues other creative endeavors, specifically photography and writing, both fiction and poetry. He has submitted his pictures to exhibits and was an active contributor to the storytelling website Cowbird until it shuttered in 2017. As a testament to his willingness to embrace serendipity, when the website’s founder held a farewell party for Cowbird last summer in Vermont, Dr. Frank drove up to celebrate with 30 or so people, most of whom were strangers to each other.
He is also, he jokes, a budding inventor, having designed a fine-dining implement called a “sporknik”—fork on one end, knife on the other, and spoon in the middle—which his son-in-law 3-D printed in stainless steel. (Buy yours online for $130.) “Joachim can get so excited about something, like a kid,” says Dr. Hashem, who was a postdoc in the lab between 2010 and 2014. “Whether it’s a new theory or a scientific article he read, or something in the newspaper, or anything at all, he would just come spill it out and get us talking about it.”
Although Dr. Frank rarely brought these parallel pursuits into the lab, his group was well aware of their place in his life. And of course, they contributed to the scientific work in their own ways, most directly in the writing: “You could give him a draft of a paper and then he changed something here and there, and suddenly it was a golden text,” says Dr. Spahn. But also in scientific imaging. Soon after Dr. Agrawal, now a senior scientist at the Wadsworth Center and a professor at the University at Albany, struck out on his own, he emailed Dr. Frank a paper describing one of the first structures his lab solved. “Within half an hour he wrote me back about the figure, and said, ‘Raj I think the colors are biting each other, you have to change it,’” Dr. Agrawal says. “He was very, very particular about first impressions of the pictures.”
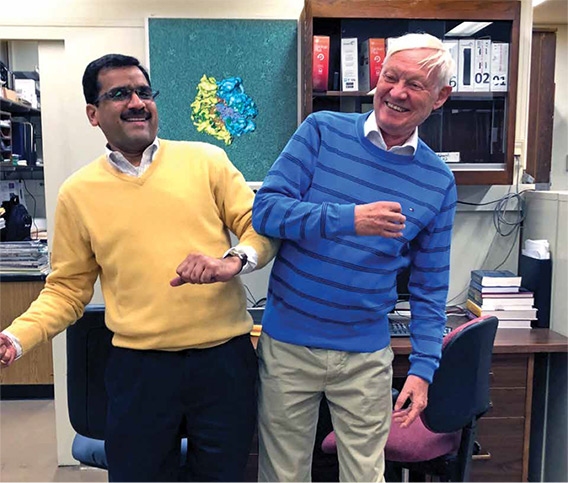 Dr. Agrawal and Dr. Frank frequently dress in the colors of the ribosome’s subunits when they get together. This photo—with the ribosome depicted in the background—was taken when Dr. Frank recently visited Dr. Agrawal’s lab at the Wadsworth Center.
Dr. Agrawal and Dr. Frank frequently dress in the colors of the ribosome’s subunits when they get together. This photo—with the ribosome depicted in the background—was taken when Dr. Frank recently visited Dr. Agrawal’s lab at the Wadsworth Center.Indeed, color choices in paper figures have long been a standing joke between Dr. Frank and Dr. Agrawal. When the two researchers began to make serious progress on the ribosome in the mid-1990s, Dr. Frank established the colors of the two subunits: light blue for the larger subunit, pale yellow for the smaller one. One morning in 2005, while attending the same conference, they emerged from nearby guesthouses with great amusement to find each other wearing sweaters of the complementary hue: Dr. Frank in light blue and Dr. Agrawal in pale yellow. “So, here we are, the very small and large subunits, and we somehow started to dance, moving our hands like the conformational change between the two subunits that we had discovered!”
Dr. Frank is also known for putting his writing skills and his penchant for the absurd on display at an annual cryo-EM conference, which generally ends with an absurdist skit. Each year he finds himself with a central role in creating or performing the skits. One year he and his colleagues transformed the trailer of the 2009 science fiction movie “Avatar” by replacing the actors’ faces with those of key researchers in the community and changing the dialogue to frets about achieving better resolution. “Every time, it was extremely hilarious,” says Dr. Hashem. “This is really where you see his great sense of humor.”
The Early Morning Nobel Call
Dr. Frank was fast asleep when the Nobel announcement was made in the wee hours of Oct. 4. He heard the telephone ring around 5:20 a.m. but attributed the commotion to a new puppy yapping to be taken outside. About 150 miles due north, in Albany, Dr. Agrawal was up early to watch, as he has for the past few years, the Nobel Foundation’s live telecasts, sure that he would eventually hear his old mentor’s name. This year it finally happened: “Cool chemistry! He said cool chemistry!” Dr. Agrawal whooped excitedly to his wife, Dr. Manjuli Sharma. “This is Joachim—it can’t be anyone else!”
Dr. Frank invited Dr. Agrawal, Dr. Spahn, Dr. Carazo, Dr. Hashem, and Mr. Grassucci, along with their families, to join him and his family in Stockholm for the week of Nobel festivities in December. “That week I saw him the happiest Joachim I have ever seen him,” says Dr. Spahn. Dr. Frank describes the experience as “a fairy tale.” On a night off from the formal events of the week, his colleagues—his scientific offspring, as it were—presented him with a gift: a telescope. “It represents a different direction of viewing the world from the one I was familiar with,” he wrote on his blog.
Not that the familiar direction has lost its allure: As this new era of cryo-EM unfolds, much exciting work remains to be done. “The enthusiasm from the recent breakthroughs is completely justified, but it is still not always so easy to get high-resolution structures,” says Dr. Spahn. “All of this is at the border of chemistry and biology, where randomly colliding molecules start to have ordered features. This is where biological complexity is created.”
Dr. Frank is particularly excited about the prospect of using cryo-EM to track how molecules within the cell change their shape to do their jobs. His group is working with a lab in Wisconsin to map this dynamic process, the so-called conformational continuum of molecules. Traditionally, researchers have assumed that molecules going from one conformation to another do so by taking the lowest-energy route, but that turns out not to be the case, he says. Instead, they can pass through many states that are energetically counterintuitive yet still necessary to their function. “You can really see how molecules actually behave,” he says.
As Dr. Frank forges ahead, Columbia University too has expanded its support for cryo-EM. Plans call for building an entire core facility dedicated to the technique. That means that after sharing a couple of microscopes and other equipment with multiple labs for the past few years, the group will soon have access to new state-of-the-art digs with three new high-end machines. “The sky is wide open now,” Dr. Frank says.
- Log in to post comments


